NAPPA Technology
 Protein microarrays display proteins in high spatial density on a microscopic surface. They can be used to test a variety of functions of many proteins simultaneously including interactions with other macromolecules, functional activity, and their suitability as substrates of enzymes. Protein microarrays are not yet widely used. In part, this may be due to the cost and technical issues associated with the expression
and purification of thousands of different proteins for printing on microarrays using traditional methods. Historically, these arrays have been produced by expressing and purifying proteins, which are then printed on the array surface. However, many challenges accompany this approach including the difficulty in purifying many proteins, a dynamic range in yields that spans several orders of magnitude (which is reflected in the widely varying amount of protein displayed on the array surface) and the danger or proteins becoming unfolded during the many manipulations, such as purification, storage and printing.
Protein microarrays display proteins in high spatial density on a microscopic surface. They can be used to test a variety of functions of many proteins simultaneously including interactions with other macromolecules, functional activity, and their suitability as substrates of enzymes. Protein microarrays are not yet widely used. In part, this may be due to the cost and technical issues associated with the expression
and purification of thousands of different proteins for printing on microarrays using traditional methods. Historically, these arrays have been produced by expressing and purifying proteins, which are then printed on the array surface. However, many challenges accompany this approach including the difficulty in purifying many proteins, a dynamic range in yields that spans several orders of magnitude (which is reflected in the widely varying amount of protein displayed on the array surface) and the danger or proteins becoming unfolded during the many manipulations, such as purification, storage and printing.
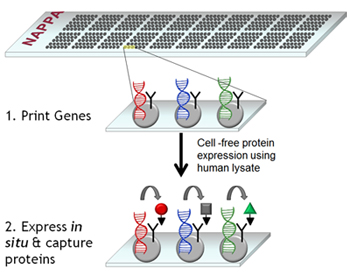
To avoid these challenges and to produce the most functional arrays possible, we have developed a novel protein microarray technology, called Nucleic Acid-Programmable Protein Array (NAPPA), which replaces the complex process of spotting purified proteins with the simple process of spotting plasmid DNA. Our technology exploits the ability to transfer protein encoding regions (open reading frames; ORFs) into specialized tagged expression vectors (all plasmids can be found through our DNASU Plasmid Repository). Proteins corresponding to the cDNAs are produced in situ as needed at the time of the assay by human in vitro transcription and translation (IVTT)-coupled cell lysates. The cDNAs are configured to append a common epitope tag to all of the proteins on their C-termini so that they can be captured by a high-affinity capture reagent that is immobilized along with the cDNA (see image above).
This method minimizes direct manipulation of the proteins and produces them just-in-time for the experiment, avoiding problems with protein purification and stability (Science. 2004 305:86 and Nat Methods. 2008 5:535) This approach offers the following advantages over conventional methods:
- Replaces printing proteins with the more reliable and less expensive process of printing DNA.
- Avoids the need to express, purify and store proteins
- Displays better than 95% of sequence-verified full-length proteins. We have demonstrated its utility in assessing sero-reactivity to membrane proteins on NAPPA;
- Protein display levels are consistent from protein to protein with 93% of proteins within two fold of the mean.
- Assures protein integrity by using both mammalian expression machinery and chaperone proteins to synthesize and fold proteins
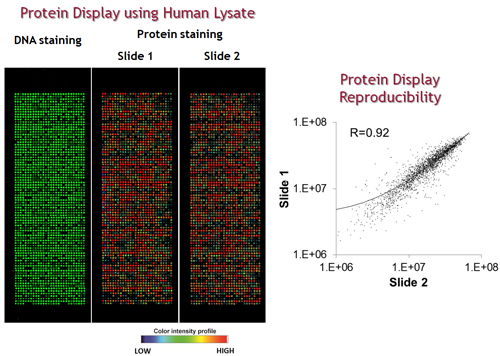
An example of a typical NAPPA array is shown below. DNA is visualized by PicoGreen and protein display by probing with an anti-GST antibody).The power of this approach is that by expressing many proteins on a single array, it is possible to test the function of many proteins simultaneously.
Video
We have a video showing each step of the NAPPA production and expression process. This does not include DNA purification using the DNA factory or serum screening using the HSPro; however, it will provide you with a good general idea of how the NAPPA process works. To view these videos, you can go to this page or visit our YouTube site.
Biological Applications
In our laboratory we use NAPPA technology to explore several biological questions including:
- Serum screening to identify antibody immune responses for a variety of diseases including cancer, diabetes, autism and infectious diseases
- Detect post-translational modifications such as phosphorylation, AMPylation, or citrullination
- Identify protein-protein or protein-DNA interactions
- Quantify protein binding kinetics in high throughput by coupling NAPPA with surface plasmon resonance (SPRi)
Click on the links above for selected images and descriptions of biological applications. For more details you can peruse our publications or visit the Center for Personalized Diagnostics website.

